Patented Anti-Correlation Technology for Capillary (Sanger) Sequencing Analysis
Mutation Surveyor software includes patented anti-correlation technology, which physically compares sample sequence traces to a reference trace, providing accuracy up to 99.5% with Phred 20 bi-directional sequences and a sensitivity of 5% of the primary peak.
Mutation Surveyor® software features patented physical comparison “anti-correlation” (U.S. Patent 8,086,410) technology and a unique multi-step alignment algorithm to rapidly and accurately detect DNA variants from Sanger Sequencing traces, including single nucleotide polymorphisms (SNPs), insertions and deletions (indels), duplications, and low frequency variants associated with mosaicism or heteroplasmy. Compatible with outputs from all major sequencing platforms, Mutation Surveyor software provides an unattended operation that is capable of detecting variants in 2000 sequence traces in less than 15 minutes. Accuracy is greater than 99% using Phred 20 bi-directional sequence traces, and sensitivity greater than 5% of the primary peak, or 1 part diseased in 17 parts normal cells. The software's Windows® based operation features a biologist-friendly interface, automatic download of GenBank reference files (GRCh37 or GRCh38) for complete annotation, contig assembly, and custom reporting that is superior in nature to those features found in similar programs such as Thermo Fisher Scientific SeqScape®, Variant Reporter™, and Minor Variant Finder; DNAStar's SeqMan Pro™; Gene Code's Sequencher®; or JSI Medical Systems' SEQUENCE PILOT.
Anti-Correlation Technology Provides Variant Accuracy >99%
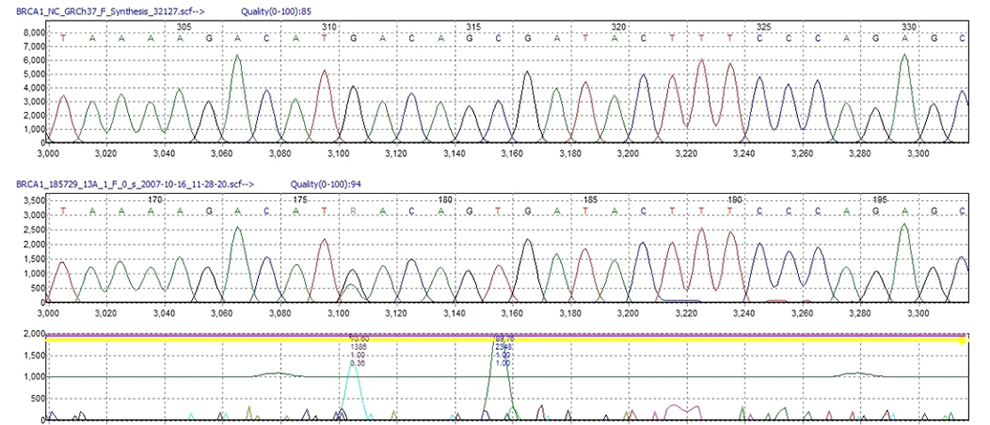
Figure 1: Two Single Nucleotide Polymorphisms detected in a sample trace. The anti-correlation technology of Mutation Surveyor software displays peaks in the “Mutation Electropherogram” (also known as Mutation Trace) where the sample peak profile differs from the reference peak profile.
Accuracy of the software in the bi-directional analysis mode is over 99% with Phred-20 sequence traces, and our collaborators have demonstrated an accuracy of 95% when processing single-directional sequence traces. The software's patented anti-correlation technology compares the sample trace to the reference trace at each nucleotide position, using physical changes in nucleotide peak profiles as a basis for SNP detection to provide excellent accuracy and sensitivity. Changes in sample traces relative to the reference are determined through three physical characteristics of the sample trace: dropping factor, overlapping factor, and signal-to-noise ratio. Peak differences are displayed in the “Mutation Electropherogram” in the middle of the screen, with larger peaks indicating greater discordance between the sample and reference position. Each mutation is given a Phred-like confidence score calculated from the physical differences of the sample and reference, providing the user with a confidence value for each mutation call.
Robust Alignment Algorithm Accurately Detects Homozygous and Heterozygous Insertions, Deletions, and Duplications
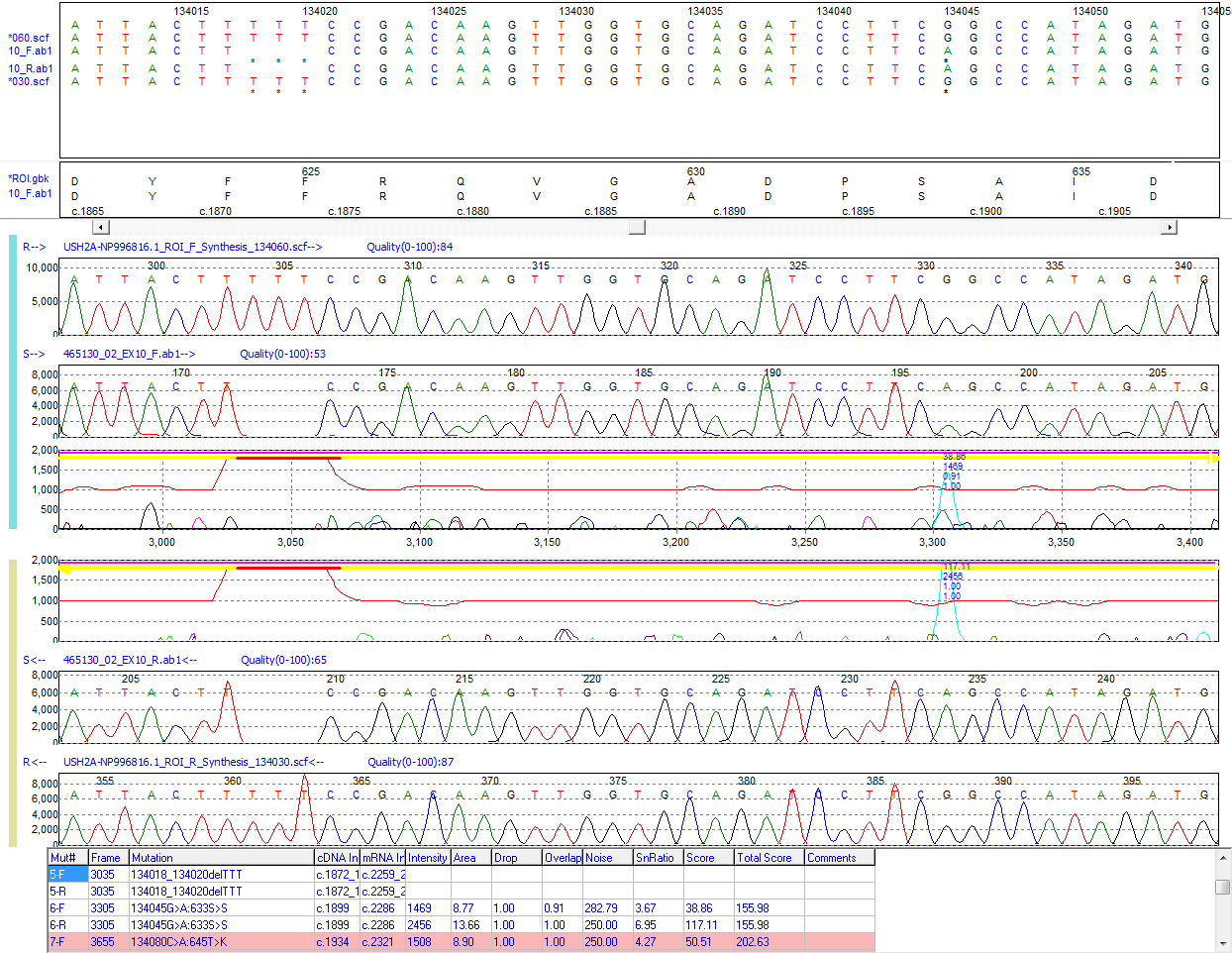
Figure 2: A homozygous deletion (TTT) and a SNP detected in a sample trace. Mutation Surveyor software’s unique multi-step alignment algorithm gaps the sample trace at the position of the deletion (red bar) and continues sample alignment to the reference trace.
Mutation Surveyor software also features a unique multi-step alignment algorithm for alignment of traces and insertion and deletion (indel) detection. While other software packages rely on base calls to determine indels, Mutation Surveyor software remains insensitive to base call errors by monitoring DNA sample migration time during sample trace alignment to the reference trace. Sample migration time is then contracted (insertion) or expanded (deletion) to accurately align sample traces to the reference to detect and display indel events.
Sensitivity Down to 5% Primary Peak
Sensitivity is greater than 5% of the primary peak, which allows detection of variants buried in the baseline of both the forward and reverse traces, alerting researchers to the presence of possible low frequency mutations.
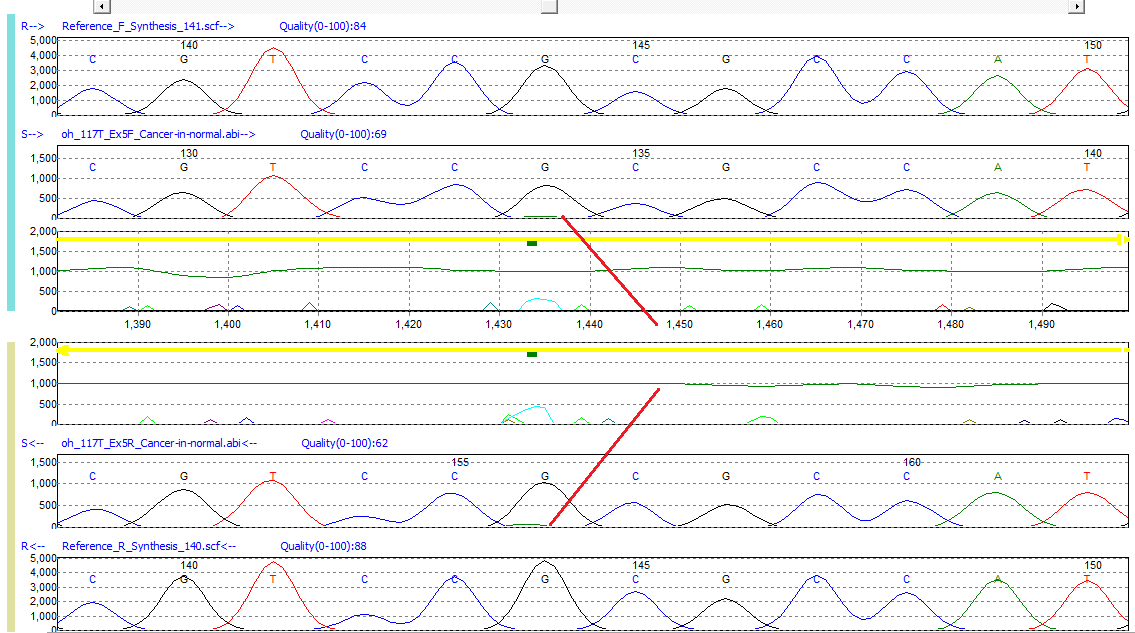
Figure 3: Somatic mutation G>GA detected in TP53. The sensitivity of Mutation Surveyor software can detect minor alleles that are present within the baseline of bi-directionally aligned peaks. Sensitivity is greater than 5% of the primary peak.
BasePatch Function Corrects Sequencing Errors
BasePatch is a function unique to Mutation Surveyor software and is not present in any other software package. This function helps to reduce baseline noise, sharpen peaks, and correct for mobility shift problems in sequence traces. When used in conjunction with our anti-correlation algorithm, it will resolve "N" calls made by the original basecall software, reduce false positives due to baseline noise, and align the ends of traces more efficiently for an extended region of mutation detection.
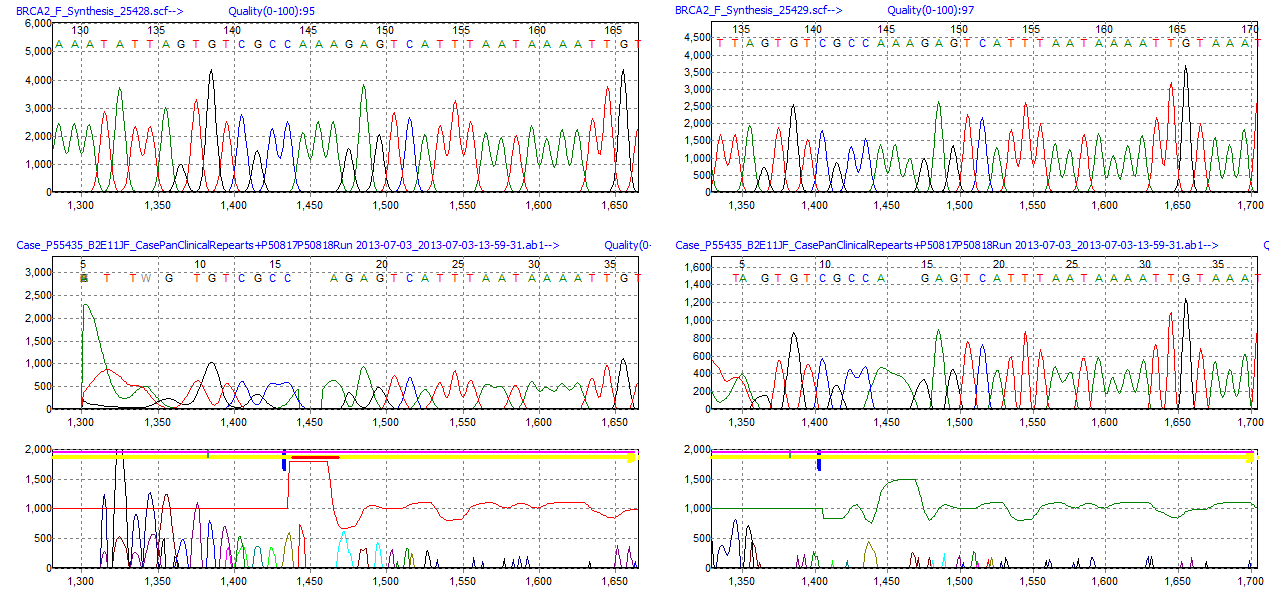
Figure 4: This illustration shows two analyses with the same parameters except that the right image has BasePatch activated while the left image does not. You can see that the trace on the right has less noise in the Mutation Electropherogram than the image on the left. Also, the lane quality score and Phred-like score increased for the analysis with BasePatch on. BasePatch is intended to increase the overall quality of the trace for better alignment and mutation detection.
Webinars
- Mutation Surveyor Software Introductory Overview
- Working With Your Mutation Surveyor Project in the Graphical Analysis Display Part I
- Working With Your Mutation Surveyor Project in the Graphical Analysis Display Part II
- Working With Your Mutation Surveyor Project in the Graphical Analysis Display Part III
- Optimizing Analysis Settings with Mutation Surveyor- Part 1
- Optimizing Analysis Settings with Mutation Surveyor- Part 2
Trademarks property of their respective owner













