Methylation Detection from Bisulfite-Treated Sanger Sequencing Traces
Methylation of DNA at cytosine positions of CpG dinucleotides has been closely associated with gene regulation, including genomic imprinting, differential gene expression, and human disease. Bisulfite treatment of DNA followed by sequencing is one way to study DNA methylation, which converts unmethylated cytosine positions to uracil, leaving methylated positions unaffected. Following PCR amplification and nucleotide sequencing, cytosine positions that are not methylated will be replaced with thymine and methylated positions will remain as cytosine. While many software packages struggle to align and detect methylation in bisulfite-treated DNA, the process is simplified in Mutation Surveyor software through a unique methylation application. Users simply load a GenBank reference file and the software will automatically convert cytosine positions to thymine nucleotides in the reference. Mutation Surveyor software quickly and accurately aligns the bisulfite-treated samples to this modified reference trace, automatically detecting methylated positions through its patented trace-to-trace “anti-correlation” technology.
Alignment and Detection of Methylated Traces
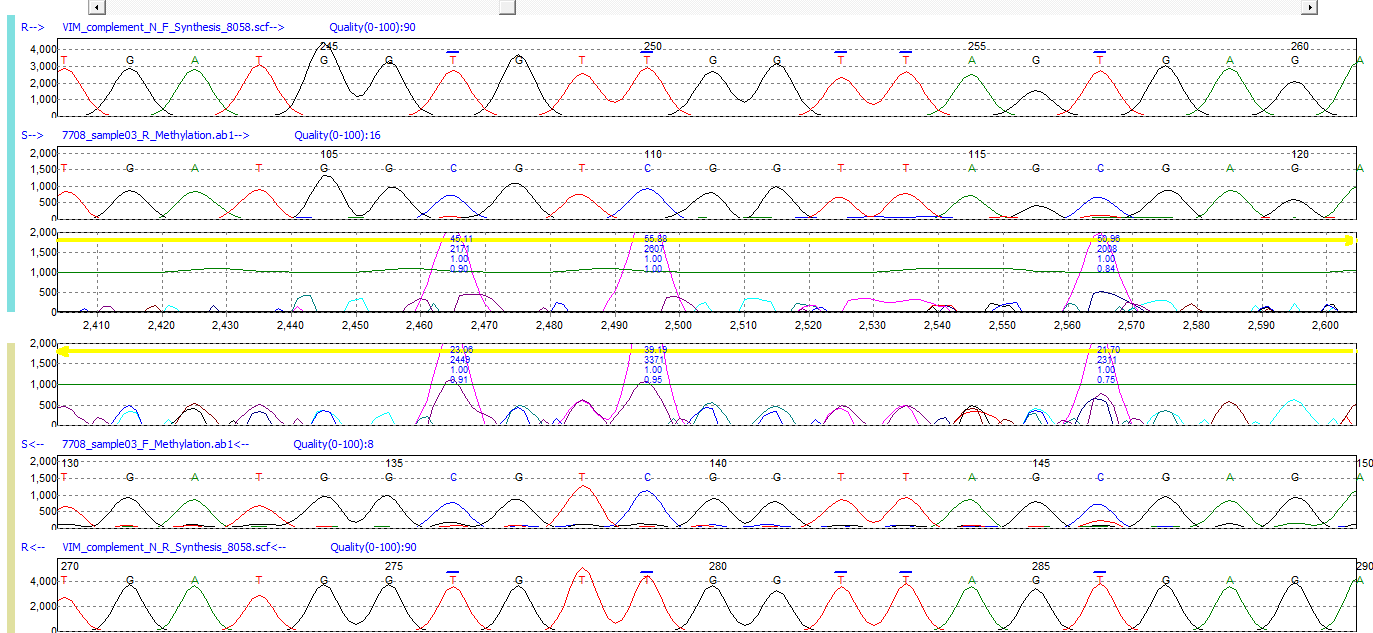
Figure 1: Mutation Surveyor software is able to accurately detect methylation in bisulfite-treated sample sequences by modifying the nucleotide sequence of the reference file. With the methylation option selected, Mutation Surveyor will convert cytosine positions to thymine in the reference file. Methylated cytosine positions in the sample trace are then detected through Mutation Surveyor software's physical trace-to-trace comparison, which displays methylated cytosine positions as spikes in the Mutation Electropherogram.
Methylation Report Provides Methylation Status of Cytosine Positions
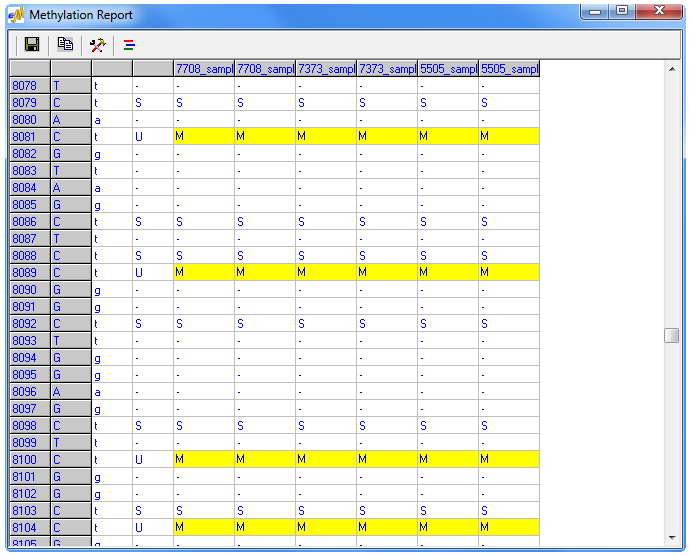
Figure 2: The Methylation report of Mutation Surveyor automatically determines the methylation status of cytosine nucleotides in the sample traces. Cytosine positions within CpG islands are labelled as methylated (M) or unmethylated (U). Cytosine positions outside of CpG islands are labelled as an incomplete (I) conversion to thymine or successful (S) conversion to thymine. Minimum methylation thresholds based on the peak profiles can be adjusted in the settings option of the report to distinguish between methylated and non-methylated cytosines.
Methylation Graphic Display Report Highlights Methylation Patterns
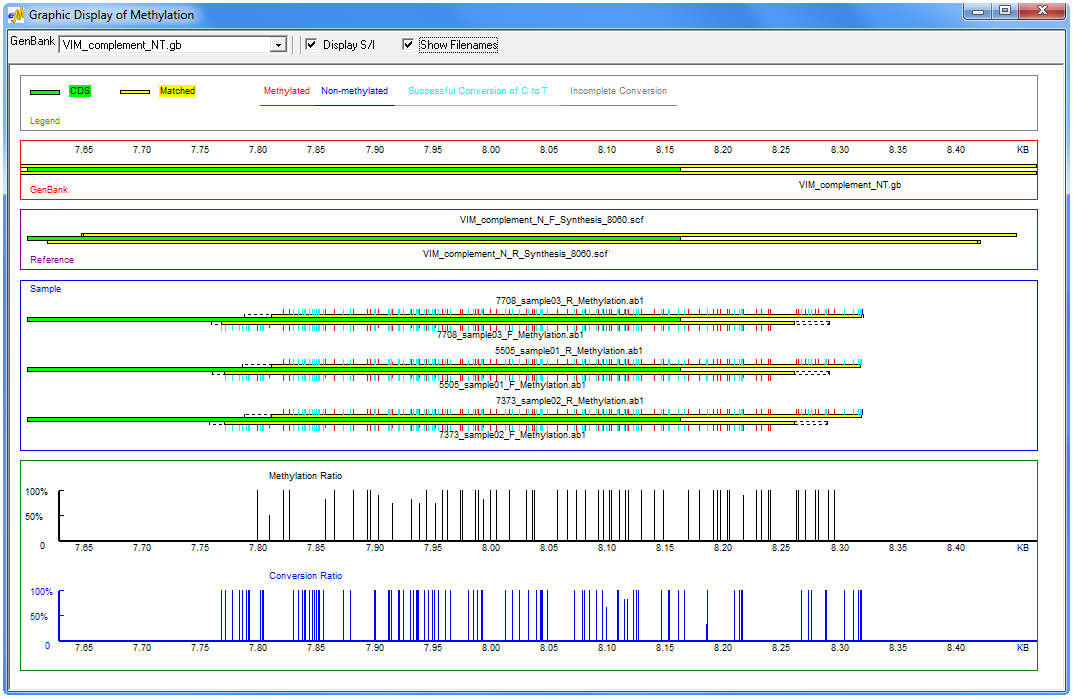
Figure 3: The Graphic Display Report of the methylation application provides a graphical representation of sample file alignment to the GenBank reference and a representation of the methylation pattern for all samples in a mutation project. Red tick marks represent methylated cytosines within CpG islands and light blue tick marks represent non-methylated cytosines outside of CpG islands. The bottom of the report provides the relative conversion ratio for every cytosine position in the project.
Application Notes:
Webinars:
- Optimizing Analysis Settings with Mutation Surveyor- Part 2
- Methylation Sequence Analysis Using Mutation Surveyor Software
Trademarks property of their respective owner













